Learning Objectives
Learning Objectives
In this section, you will explore the following questions:
- What are different types of mutations in DNA and the significance of mutations?
- What are examples of mechanisms that repair mutations in DNA?
Connection for AP® Courses
Connection for AP® Courses
DNA polymerase is an efficient enzyme but it can make mistakes while adding nucleotides during replication. It edits the DNA by proofreading every newly added base. An incorrect base is removed and replaced by the correct base. If a base remains mismatched, special repair enzymes can often recognize the wrongly incorporated base, excise it from the DNA, and replace it with the correct base. Most mistakes are corrected, but if they are not they may result in a mutation, which is defined as a permanent change in a DNA sequence. A mutation can be passed to daughter cells through DNA replication and cell division. There are several types of DNA mutations, including substitution, deletion, insertion, and translocation. Mutations in repair genes may lead to serious consequences, such as cancer. Mutations can be induced by environmental factors, such as UV radiation, or they can occur spontaneously. We will explore the effects of mutation in more detail in a later chapter. Remember that mutations are not always detrimental. They can be beneficial, too. Changes in DNA increase genetic variation—the foundation of evolution by natural selection.
Information presented and the examples highlighted in the section support concepts outlined in Big Idea 3 of the AP® Biology Curriculum Framework. The Learning Objectives listed in the Curriculum Framework provide a transparent foundation for the AP® Biology course, an inquiry-based laboratory experience, instructional activities, and AP® exam questions. A Learning Objective merges required content with one or more of the seven Science Practices.
Big Idea 3 Living systems store, retrieve, transmit, and respond to information essential to life processes.
Enduring Understanding 3.C The processing of genetic information is imperfect and is a source of genetic variation.
| Essential Knowledge | 3.C.1 Changes in genotype can result in changes in phenotype. |
| Science Practice | 6.4 The student can make claims and predictions about natural phenomena based on scientific theories and models. |
| Science Practice | 7.2 The student can connect concepts in and across domain(s) to generalize or extrapolate in and/or across enduring understandings and/or big ideas. |
| Learning Objective | 3.24 The student is able to predict how a change in genotype, when expressed as a phenotype, provides a variation that can be subject to natural selection. |
| Essential Knowledge | 3.C.1 Changes in genotype can result in changes in phenotype. |
| Science Practice | 1.1 The student can create representations and models of natural or man-made phenomena and systems in the domain. |
| Learning Objective | 3.25 The student can create a visual representation to illustrate how changes in a DNA nucleotide sequence can result in a change in the polypeptide produced. |
| Essential Knowledge | 3.C.2 Biological systems have multiple processes that increase genetic variation. |
| Science Practice | 6.2 The student can construct explanations of phenomena based on evidence produced through scientific practices. |
| Learning Objective | 3.28 The student is able to construct an explanation of the multiple processes that increase variation within a population. |
DNA replication is a highly accurate process, but mistakes can occasionally occur, such as a DNA polymerase inserting a wrong base. Uncorrected mistakes may sometimes lead to serious consequences, such as cancer. Repair mechanisms correct the mistakes. In rare cases, mistakes are not corrected, leading to mutations; in other cases, repair enzymes are themselves mutated or defective.
Most of the mistakes during DNA replication are promptly corrected by DNA polymerase by proofreading the base that has been just added (Figure 14.17). In proofreading, the DNA pol reads the newly added base before adding the next one, so a correction can be made. The polymerase checks whether the newly added base has paired correctly with the base in the template strand. If it is the right base, the next nucleotide is added. If an incorrect base has been added, the enzyme makes a cut at the phosphodiester bond and releases the wrong nucleotide. This is performed by the exonuclease action of DNA pol III. Once the incorrect nucleotide has been removed, a new one will be added again.
Some errors are not corrected during replication, but are instead corrected after replication is completed; this type of repair is known as mismatch repair (Figure 14.18). The enzymes recognize the incorrectly added nucleotide and excise it; this is then replaced by the correct base. If this remains uncorrected, it may lead to more permanent damage. How do mismatch repair enzymes recognize which of the two bases is the incorrect one? In E. coli, after replication, the nitrogenous base adenine acquires a methyl group; the parental DNA strand will have methyl groups, whereas the newly synthesized strand lacks them. Thus, DNA polymerase is able to remove the wrongly incorporated bases from the newly synthesized, non-methylated strand. In eukaryotes, the mechanism is not very well understood, but it is believed to involve recognition of unsealed nicks in the new strand, as well as a short-term continuing association of some of the replication proteins with the new daughter strand after replication has completed.
In another type of repair mechanism, nucleotide excision repair, enzymes replace incorrect bases by making a cut on both the 3' and 5' ends of the incorrect base (Figure 14.19). The segment of DNA is removed and replaced with the correctly paired nucleotides by the action of DNA pol. Once the bases are filled in, the remaining gap is sealed with a phosphodiester linkage catalyzed by DNA ligase. This repair mechanism is often employed when UV exposure causes the formation of pyrimidine dimers.
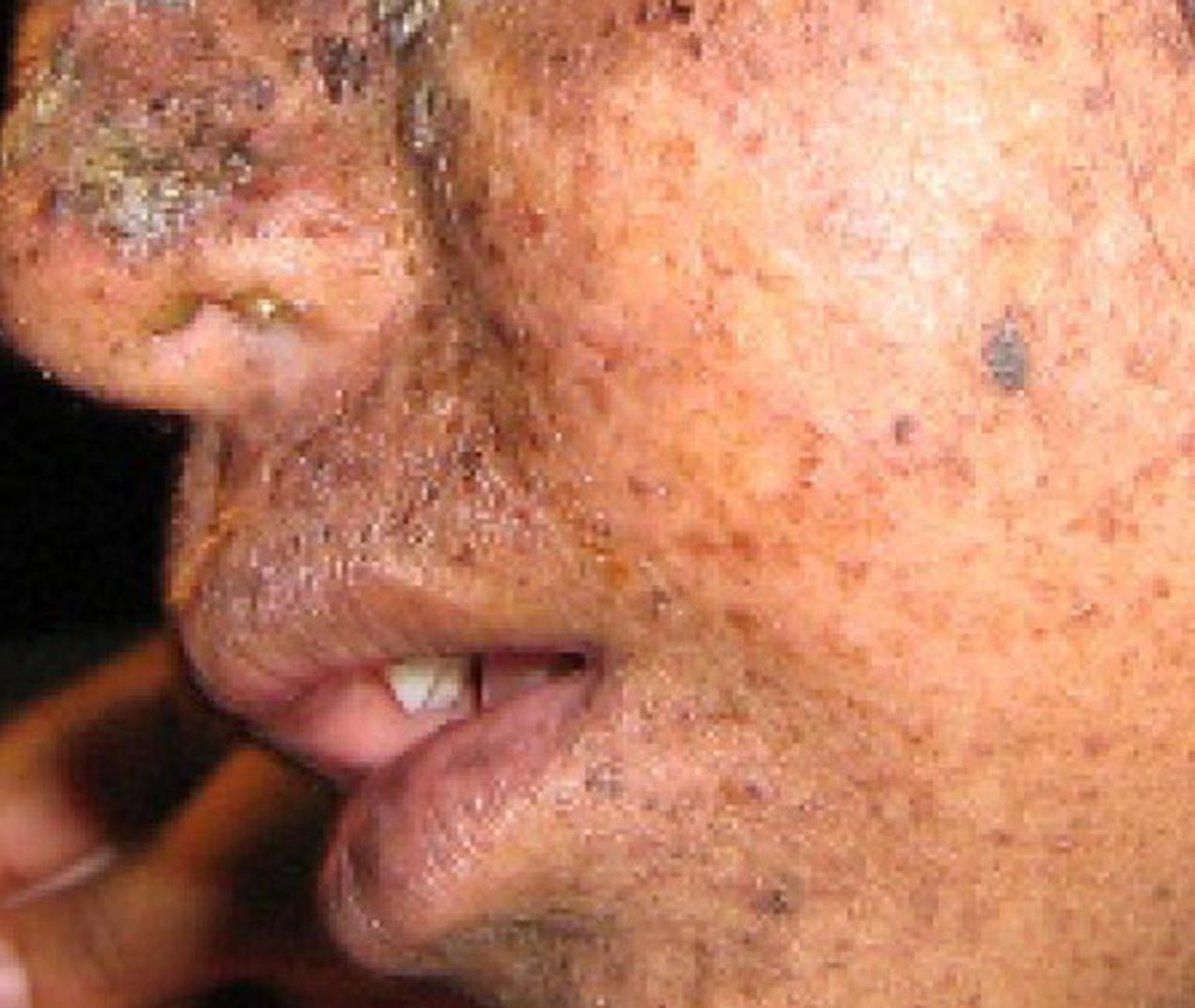
A well-studied example of mistakes not being corrected is seen in people suffering from xeroderma pigmentosa (Figure 14.20). Affected individuals have skin that is highly sensitive to UV rays from the sun. When individuals are exposed to UV, pyrimidine dimers, especially those of thymine, are formed; people with xeroderma pigmentosa are not able to repair the damage. These are not repaired because of a defect in the nucleotide excision repair enzymes, whereas in normal individuals, the thymine dimers are excised and the defect is corrected. The thymine dimers distort the structure of the DNA double helix, and this may cause problems during DNA replication.
Errors during DNA replication are not the only reason why mutations arise in DNA. Mutations, variations in the nucleotide sequence of a genome, can also occur because of damage to DNA. Such mutations may be of two types: induced or spontaneous. Induced mutations are those that result from an exposure to chemicals, UV rays, X-rays, or some other environmental agent. Spontaneous mutations occur without any exposure to any environmental agent; they are a result of natural reactions taking place within the body.
Mutations may have a wide range of effects. Some mutations are not expressed; these are known as silent mutations. Point mutations are those mutations that affect a single base pair. The most common nucleotide mutations are substitutions, in which one base is replaced by another. These can be of two types, either transitions or transversions. Transition substitution refers to a purine or pyrimidine being replaced by a base of the same kind; for example, a purine such as adenine may be replaced by the purine guanine. Transversion substitution refers to a purine being replaced by a pyrimidine, or vice versa; for example, cytosine, a pyrimidine, is replaced by adenine, a purine. Mutations can also be the result of the addition of a base, known as an insertion, or the removal of a base, also known as deletion. Sometimes a piece of DNA from one chromosome may get translocated to another chromosome or to another region of the same chromosome; this is also known as translocation. These mutation types are shown in Figure 14.22.
Everyday Connection for AP® Courses
Sometimes a nucleotide is overlooked by the DNA repair system for no known reason. This malignant melanoma is the result of DNA not undergoing repair after too much UV exposure.
Which statement about the above malignant melanoma is most likely true?
- It was the result of a spontaneous mutation.
- It was caused by thymine dimer formation.
- It was caused by a transition substitution.
- It was caused by a transversion substitution.
Visual Connection
- Addition of three nucleotides does not shift the reading frame.
- Addition of three nucleotides shifts the reading frame.
- Addition of three nucleotides incorporates two amino acids.
- Addition of three nucleotides removes two amino acids.
Mutations in repair genes have been known to cause cancer. Many mutated repair genes have been implicated in certain forms of pancreatic cancer, colon cancer, and colorectal cancer. Mutations can affect either somatic cells or germ cells. If many mutations accumulate in a somatic cell, they may lead to problems such as the uncontrolled cell division observed in cancer. If a mutation takes place in germ cells, the mutation will be passed on to the next generation, as in the case of hemophilia and xeroderma pigmentosa.
Science Practice Connection for AP® Courses
Think About It
Infertility can sometimes be explained by chromosome translocations. Explain how chromosome translocations can cause infertility. Are there times when a chromosome translocation might not result in infertility?
References
References
Jaskelioff, M. et al. (2011, Jan. 6). Telomerase reactivation reverses tissue degeneration in aged telomerase-deficient mice. Nature, 469, 102–7.