Learning Objectives
Learning Objectives
In this section, you will explore the following question:
- What is the role of transcription factors, enhancers, and repressors in gene regulation?
Connection for AP® Courses
Connection for AP® Courses
To start transcription, general transcription factors must first bind to a specific area on the DNA called the TATA box and then recruit RNA polymerase to that location. In addition, other areas on the DNA called enhancer regions help augment transcription. Transcription factors can bind to enhancer regions to increase or prevent transcription.
Information presented and the examples highlighted in the section support concepts outlined in Big Idea 3 of the AP® Biology Curriculum Framework. The Learning Objectives listed in the Curriculum Framework provide a transparent foundation for the AP® Biology course, an inquiry-based laboratory experience, instructional activities, and AP® exam questions. A Learning Objective merges required content with one or more of the seven Science Practices
| Big Idea 3 | Living systems store, retrieve, transmit, and respond to information essential to life processes. |
| Enduring Understanding 3.B | Expression of genetic information involves cellular and molecular mechanisms. |
| Essential Knowledge | 3.B.1 Gene regulation results in differential gene expression, leading to cell specialization |
| Science Practice | 7.1 The student can connect phenomena and models across spatial and temporal scales. |
| Learning Objective | 3.18 The student is able to describe the connection between the regulation of gene expression and observed differences between different kinds of organisms |
| Essential Knowledge | 3.B.1 Gene regulation results in differential gene expression, leading to cell specialization |
| Science Practice | 7.1 The student can connect phenomena and models across spatial and temporal scales |
| Learning Objective | 3.19 The student is able to describe the connection between the regulation of gene expression and observed differences between individuals in a population |
| Essential Knowledge | 3.B.1 Gene regulation results in differential gene expression, leading to cell specialization. |
| Science Practice | 6.2 The student can construct explanations of phenomena based on evidence produced through scientific practices |
| Learning Objective | 3.20 The student is able to explain how the regulation of gene expression is essential for the processes and structures that support efficient cell function. |
| Essential Knowledge | 3.B.1 1 Gene regulation results in differential gene expression, leading to cell specialization. |
| Science Practice | 1.4 The student can use representations and models to analyze situations or solve problems qualitatively and quantitatively. |
| Learning Objective | 3.21 The student can use representations to describe how gene regulation influences cell products and function. |
The Science Practices Assessment Ancillary contains additional test questions for this section that will help you prepare for the AP exam. These questions address the following standard:
- [APLO 3.18]
Like prokaryotic cells, the transcription of genes in eukaryotes requires the actions of an RNA polymerase to bind to a sequence upstream of a gene to initiate transcription. However, unlike prokaryotic cells, the eukaryotic RNA polymerase requires other proteins, or transcription factors, to facilitate transcription initiation. Transcription factors are proteins that bind to the promoter sequence and other regulatory sequences to control the transcription of the target gene. RNA polymerase by itself cannot initiate transcription in eukaryotic cells. Transcription factors must bind to the promoter region first and recruit RNA polymerase to the site for transcription to be established.
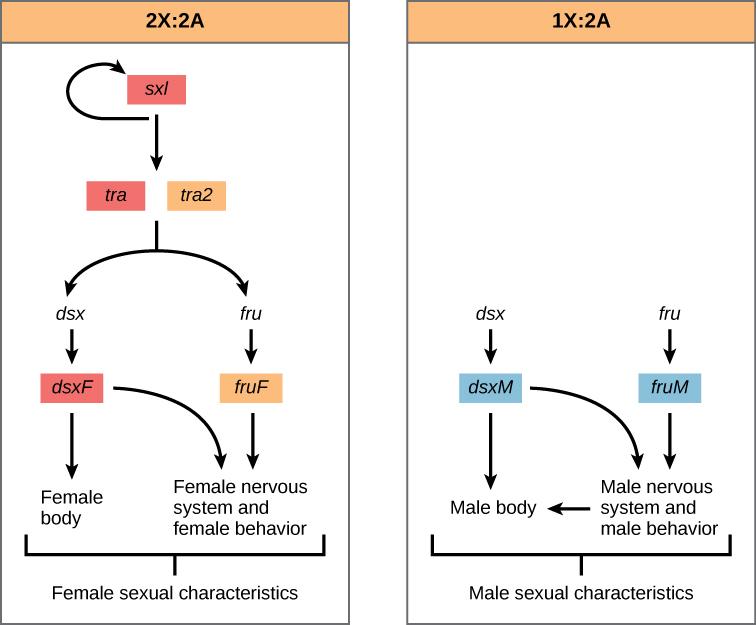
These transcriptional programs in eukaryotic organisms are responsible for the development of complex functions and behaviors. For example, the development of primary sexual characteristics is regulated by the interaction of several key genes (Figure 16.9). In Drosophila, the slx gene determines the sex. This gene is transcribed when there are two copies of the X chromosome. When it is expressed, it binds to the RNA transcript of tra and regulates its splicing. When sxl is present, the female form of tra is expressed which then binds to the transcript of dsx and fru, ultimately producing female sexual characteristics. In males, tra does not regulate the transcription of dsx and fru, meaning that default splicing occurs and different dsx and fru proteins are made, resulting in male sexual characteristics.
Link to Learning
View the process of transcription—the making of RNA from a DNA template—at this site.
- DNA unwinds, transcription factors bind, the termination complex forms, and DNA polymerase adds nucleotides to the mRNA.
- DNA unwinds, transcription factors bind, and RNA polymerase adds nucleotides to the mRNA.
- The transcription complex forms, transcription factors add nucleotides to the forming mRNA, and the mRNA disconnects from the DNA.
- Elongation occurs, followed by the formation of the transcription initiation complex and the disconnection of the mRNA strand from DNA.
The Promoter and the Transcription Machinery
The Promoter and the Transcription Machinery
Genes are organized to make the control of gene expression easier. The promoter region is immediately upstream of the coding sequence. This region can be short (only a few nucleotides in length) or quite long (hundreds of nucleotides long). The longer the promoter, the more available space for proteins to bind. This also adds more control to the transcription process. The length of the promoter is gene-specific and can differ dramatically between genes. Consequently, the level of control of gene expression can also differ quite dramatically between genes. The purpose of the promoter is to bind transcription factors that control the initiation of transcription.
Within the promoter region, just upstream of the transcriptional start site, resides the TATA box. This box is simply a repeat of thymine and adenine dinucleotides (literally, TATA repeats). RNA polymerase binds to the transcription initiation complex, allowing transcription to occur. To initiate transcription, a transcription factor (TFIID) is the first to bind to the TATA box. Binding of TFIID recruits other transcription factors, including TFIIB, TFIIE, TFIIF, and TFIIH to the TATA box. Once this complex is assembled, RNA polymerase can bind to its upstream sequence. When bound along with the transcription factors, RNA polymerase is phosphorylated. This releases part of the protein from the DNA to activate the transcription initiation complex and places RNA polymerase in the correct orientation to begin transcription; DNA-bending protein brings the enhancer, which can be quite a distance from the gene, in contact with transcription factors and mediator proteins (Figure 16.10).
In addition to the general transcription factors, other transcription factors can bind to the promoter to regulate gene transcription. These transcription factors bind to the promoters of a specific set of genes. They are not general transcription factors that bind to every promoter complex, but are recruited to a specific sequence on the promoter of a specific gene. There are hundreds of transcription factors in a cell that each bind specifically to a particular DNA sequence motif. When transcription factors bind to the promoter just upstream of the encoded gene, it is referred to as a cis-acting element, because it is on the same chromosome just next to the gene. The region that a particular transcription factor binds to is called the transcription factor-binding site. Transcription factors respond to environmental stimuli that cause the proteins to find their binding sites and initiate transcription of the gene that is needed.
Enhancers and Transcription
Enhancers and Transcription
In some eukaryotic genes, there are regions that help increase or enhance transcription. These regions, called enhancers, are not necessarily close to the genes they enhance. They can be located upstream of a gene, within the coding region of the gene, downstream of a gene, or may be thousands of nucleotides away.
Enhancer regions are binding sequences, or sites, for transcription factors. When a DNA-bending protein binds, the shape of the DNA changes (Figure 16.10). This shape change allows for the interaction of the activators bound to the enhancers with the transcription factors bound to the promoter region and the RNA polymerase. Whereas DNA is generally depicted as a straight line in two dimensions, it is actually a three-dimensional object. Therefore, a nucleotide sequence thousands of nucleotides away can fold over and interact with a specific promoter.
Turning Genes Off: Transcriptional Repressors
Turning Genes Off: Transcriptional Repressors
Like prokaryotic cells, eukaryotic cells also have mechanisms to prevent transcription. Transcriptional repressors can bind to promoter or enhancer regions and block transcription. Like the transcriptional activators, repressors respond to external stimuli to prevent the binding of activating transcription factors.
Science Practice Connection for AP® Courses
Think About It
How can cells in a multicellular eukaryotic organism be of different types given that they all share the same genome?
Disclaimer
This section may include links to websites that contain links to articles on unrelated topics. See the preface for more information.